Implementation of Wiseman many interacting worlds simulation for 1D, 1 Particle systems
###MIWsim1D###
Author: D Cyganski
Created: 2015-05-17
####Implementation of Wiseman many interacting worlds simulation for 1D, 1 Particle systems####
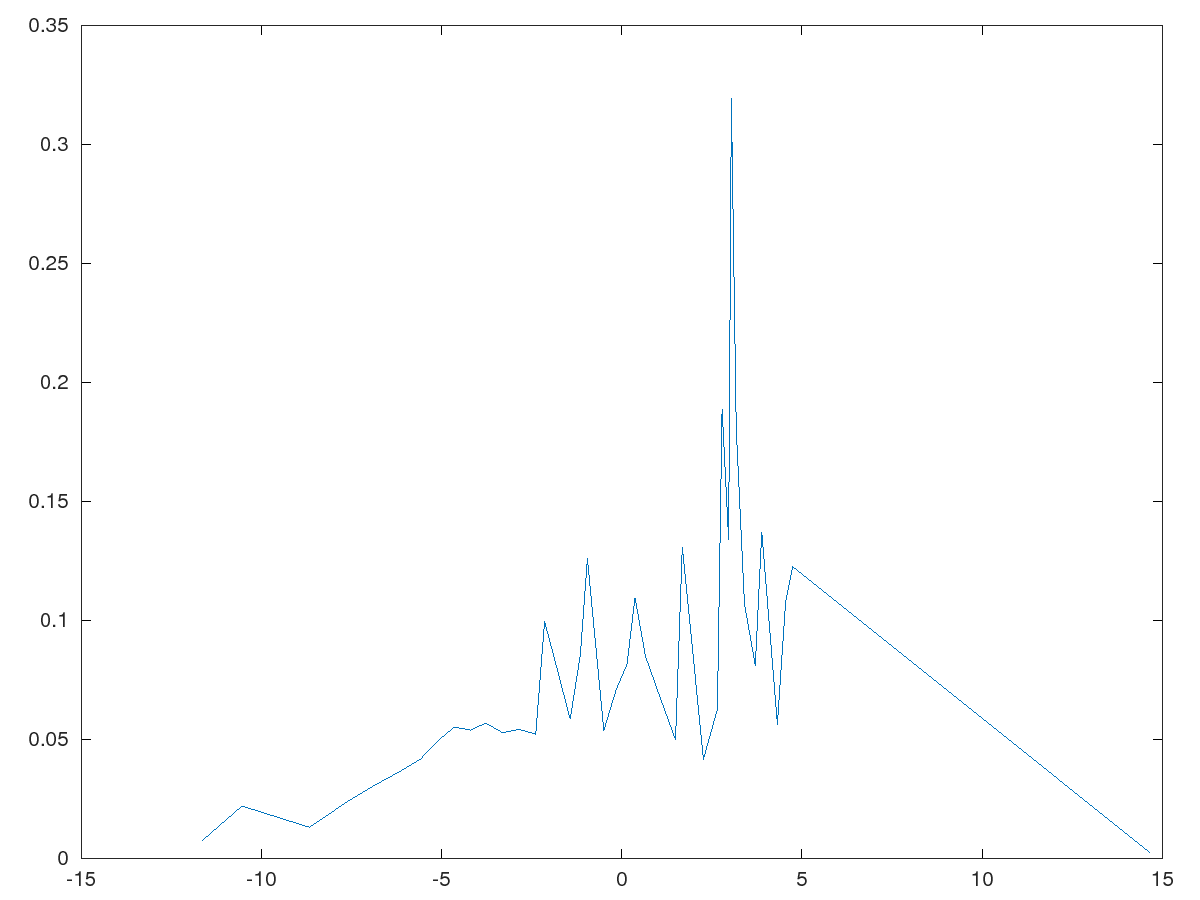
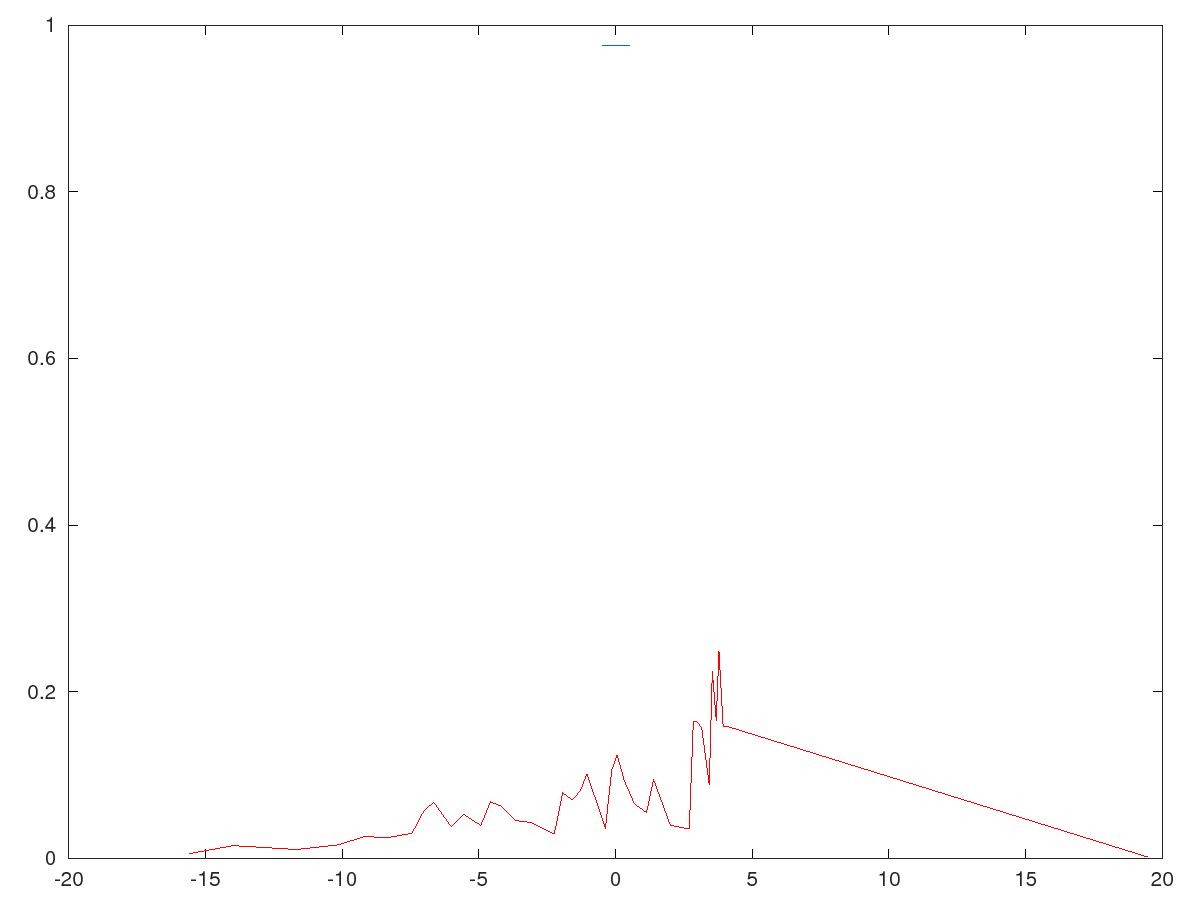
v1 - demo of packet spreading and Fresnel diffraction of 1D packet with spreading
v2 - single particle with gaussian distribution and nonzero mean velocity, colliding with a potential barrier
v3 - single particle in two gaussian superposition with velocities, colliding with no interacting fields
The future track of the many worlds position state is determined in general in MIW by system of coupled non-linear second order partial differential equations, where =number of worlds, =number of particles, =number of dimensions. where indexes the particle and the world.
Such a system of second order non-linear PDEs can always be written as a system of of first order linear PDEs:
which in turn can be put into matrix state-system form for state vector X=[x, y]'
where I have suppressed the non-linear functional dependencies of and , and incorporated the mass scaling into the new function and stacked all the particles and worlds into the single vector
We will now discretize this matrix differential equation and introduce the discrete time state vector of positions and velocities ( and ) to be named .
For our 1 D case, there will be equations. has state variables (position, velocity) In a discrete state space representation, the system is described as first order linear state transition equations, and is representable as a dimenstional matrix state space equation.

Now use the appropriate state propagation matrix to step this system forward in time. with ; where is the continuous time state propagation matrix, is the time interval between time steps, and expm is octave's matrix exponentiation function.
Note: The HDW paper does not explicitly use the matrix exponentiation based conversion of the continuous time to discrete time conversion. While a naive conversion produces results that converge to the same solution as approaches 0, for any finite value of , the exponentiation approach will produce solutions closer to that of the un-discretized system of equations.
I discovered that when I applied the exponentiation to the Ac matrix, thanks to the fact that the differential equations involved did not involve any state variables in its linear part, the outcome of expm(Ac) yields the most simple (naive) summation equation. Since the matrix A that results is extremely sparse, it is computationally advantageous to simply use direct for loop implementation than matrix multiplication.

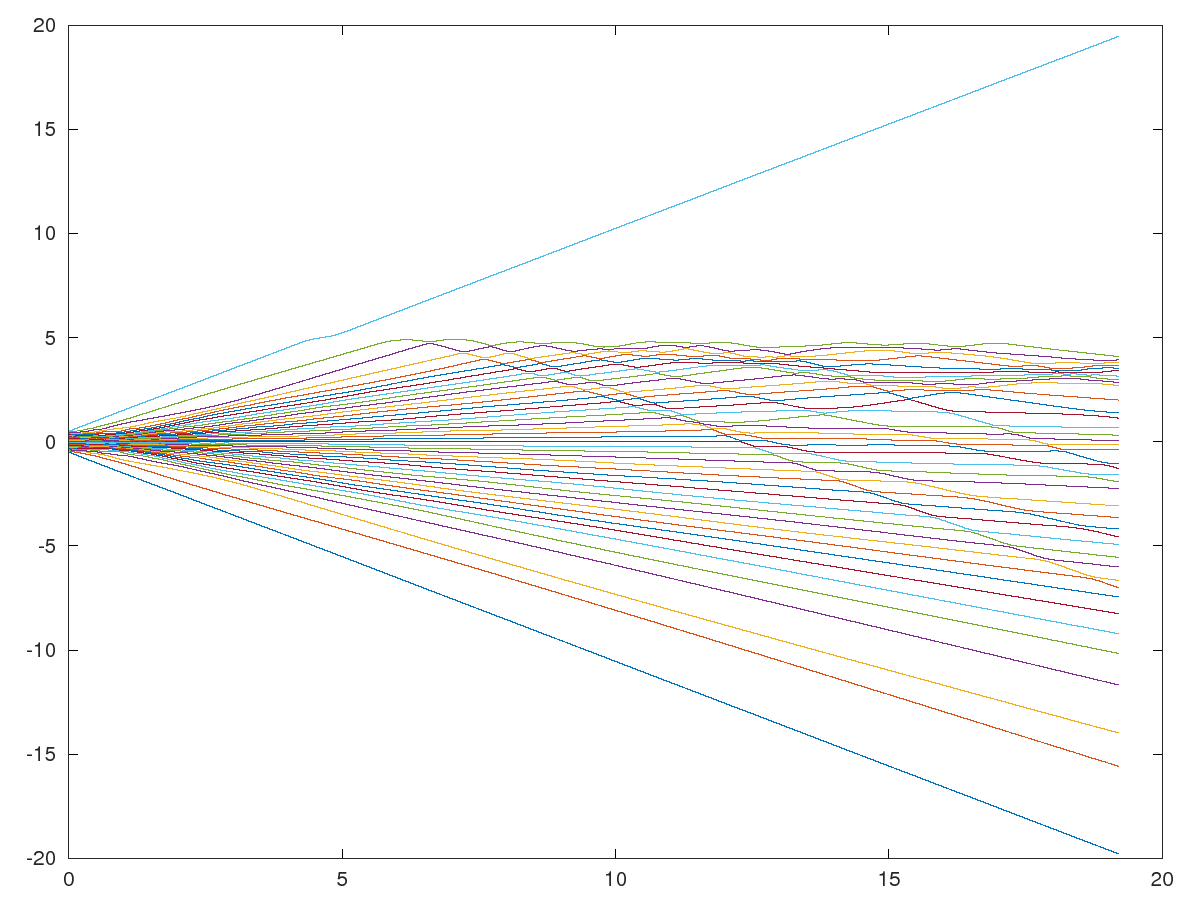

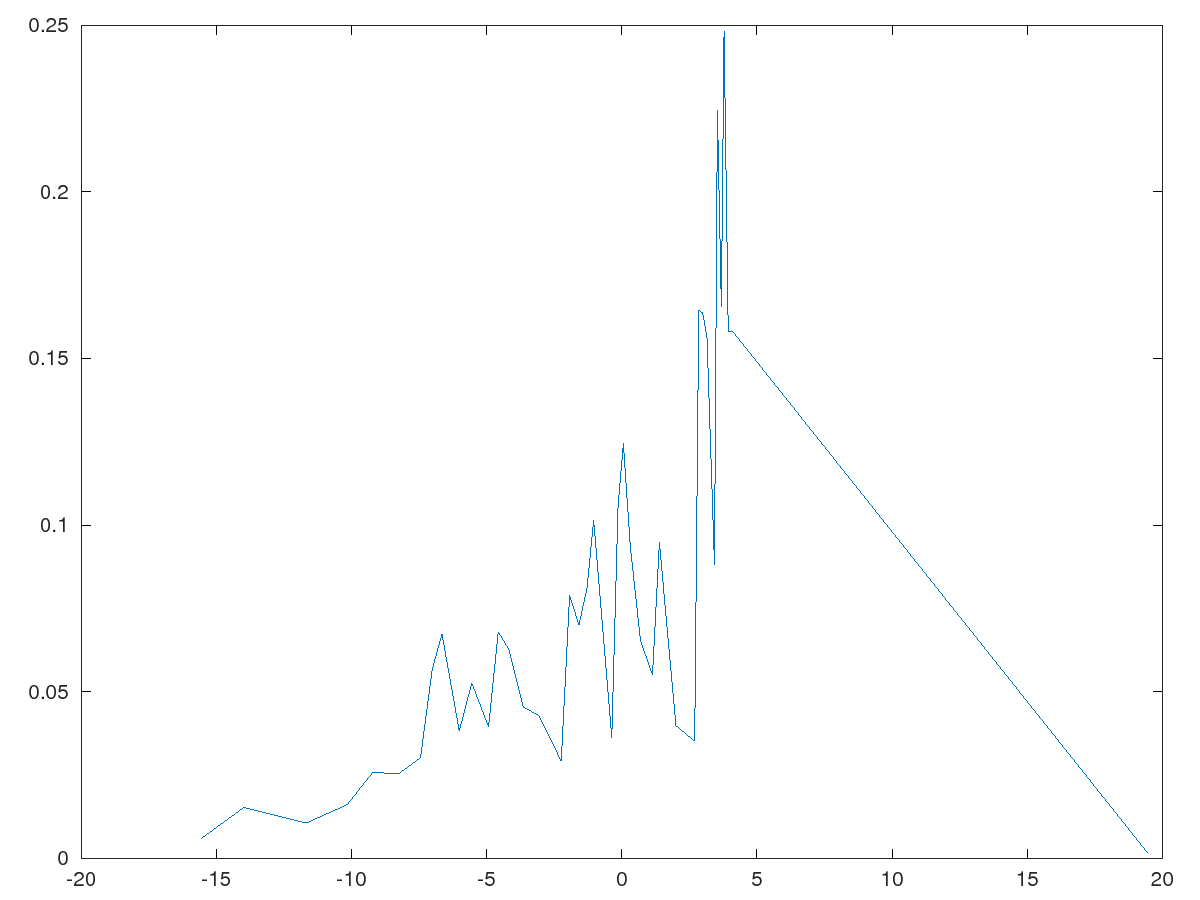
Inline plot failed, consider trying another graphics toolkit
error: print: no axes object in figure to print
error: called from
_make_figures>safe_print at line 122 column 7
_make_figures at line 44 column 13